Filter News
Area of Research
- (-) Biology and Environment (16)
- Biological Systems (1)
- Clean Energy (27)
- Computational Biology (1)
- Computational Engineering (1)
- Computer Science (6)
- Electricity and Smart Grid (1)
- Isotopes (1)
- Materials (21)
- Materials for Computing (5)
- National Security (3)
- Neutron Science (24)
- Nuclear Science and Technology (3)
- Quantum information Science (1)
- Sensors and Controls (1)
- Supercomputing (7)
News Topics
- (-) Artificial Intelligence (1)
- (-) Bioenergy (10)
- (-) Biomedical (2)
- (-) Grid (2)
- (-) Machine Learning (1)
- 3-D Printing/Advanced Manufacturing (2)
- Big Data (1)
- Biology (14)
- Biotechnology (2)
- Clean Water (3)
- Climate Change (9)
- Composites (1)
- Computer Science (3)
- Coronavirus (1)
- Decarbonization (2)
- Environment (17)
- High-Performance Computing (3)
- Hydropower (3)
- Materials (1)
- Mercury (1)
- Simulation (1)
- Sustainable Energy (9)
- Transportation (1)
Media Contacts

Researchers at Oak Ridge National Laboratory are using a novel approach in determining environmental impacts to aquatic species near hydropower facilities, potentially leading to smarter facility designs that can support electrical grid reliability.
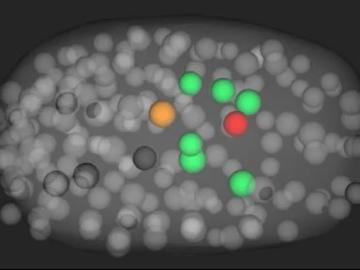
Scientists have developed a novel approach to computationally infer previously undetected behaviors within complex biological environments by analyzing live, time-lapsed images that show the positioning of embryonic cells in C. elegans, or roundworms. Their published methods could be used to reveal hidden biological activity.

An analysis by Oak Ridge National Laboratory shows that using less-profitable farmland to grow bioenergy crops such as switchgrass could fuel not only clean energy, but also gains in biodiversity.
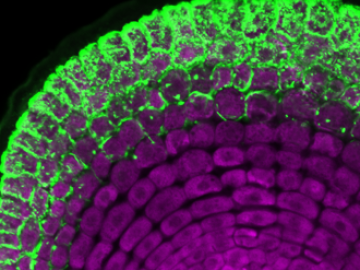
An ORNL team has successfully introduced a poplar gene into switchgrass, an important biofuel source, that allows switchgrass to interact with a beneficial fungus, ultimately boosting the grass’ growth and viability in changing environments.
Oak Ridge National Laboratory and collaborators have discovered that signaling molecules known to trigger symbiosis between plants and soil bacteria are also used by almost all fungi as chemical signals to communicate with each other.
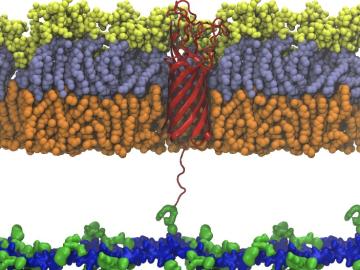
Scientists from Oak Ridge National Laboratory used high-performance computing to create protein models that helped reveal how the outer membrane is tethered to the cell membrane in certain bacteria.


