Filter News
Area of Research
News Topics
- (-) Biotechnology (22)
- (-) Isotopes (53)
- (-) Mercury (12)
- 3-D Printing/Advanced Manufacturing (122)
- Advanced Reactors (34)
- Artificial Intelligence (91)
- Big Data (55)
- Bioenergy (92)
- Biology (99)
- Biomedical (58)
- Buildings (57)
- Chemical Sciences (65)
- Clean Water (29)
- Climate Change (100)
- Composites (26)
- Computer Science (189)
- Coronavirus (46)
- Critical Materials (26)
- Cybersecurity (35)
- Decarbonization (80)
- Education (4)
- Element Discovery (1)
- Emergency (2)
- Energy Storage (109)
- Environment (195)
- Exascale Computing (37)
- Fossil Energy (6)
- Frontier (42)
- Fusion (55)
- Grid (63)
- High-Performance Computing (85)
- Hydropower (11)
- Irradiation (3)
- ITER (7)
- Machine Learning (48)
- Materials (144)
- Materials Science (141)
- Mathematics (8)
- Microelectronics (3)
- Microscopy (51)
- Molten Salt (8)
- Nanotechnology (60)
- National Security (62)
- Net Zero (14)
- Neutron Science (131)
- Nuclear Energy (109)
- Partnerships (44)
- Physics (61)
- Polymers (33)
- Quantum Computing (34)
- Quantum Science (69)
- Renewable Energy (2)
- Security (24)
- Simulation (48)
- Software (1)
- Space Exploration (25)
- Statistics (3)
- Summit (57)
- Sustainable Energy (126)
- Transformational Challenge Reactor (7)
- Transportation (97)
Media Contacts
ORNL scientists have modified a single microbe to simultaneously digest five of the most abundant components of lignocellulosic biomass, a big step forward in the development of a cost-effective biochemical conversion process to turn plants into

Radioactive isotopes power some of NASA’s best-known spacecraft. But predicting how radiation emitted from these isotopes might affect nearby materials is tricky

After its long journey to Mars beginning this summer, NASA’s Perseverance rover will be powered across the planet’s surface in part by plutonium produced at the Department of Energy’s Oak Ridge National Laboratory.

Oak Ridge National Laboratory researchers have discovered a better way to separate actinium-227, a rare isotope essential for an FDA-approved cancer treatment.
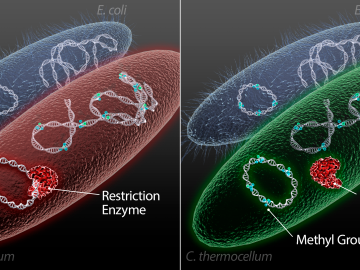
Scientists at the US Department of Energy’s Oak Ridge National Laboratory have demonstrated a method to insert genes into a variety of microorganisms that previously would not accept foreign DNA, with the goal of creating custom microbes to break down plants for bioenergy.

Sometimes solutions to the biggest problems can be found in the smallest details. The work of biochemist Alex Johs at Oak Ridge National Laboratory bears this out, as he focuses on understanding protein structures and molecular interactions to resolve complex global problems like the spread of mercury pollution in waterways and the food supply.

OAK RIDGE, Tenn., Jan. 31, 2019—A new electron microscopy technique that detects the subtle changes in the weight of proteins at the nanoscale—while keeping the sample intact—could open a new pathway for deeper, more comprehensive studies of the basic building blocks of life.
Physicists turned to the “doubly magic” tin isotope Sn-132, colliding it with a target at Oak Ridge National Laboratory to assess its properties as it lost a neutron to become Sn-131.
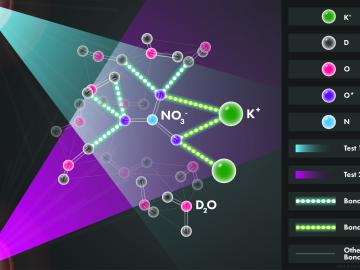
Scientists at the Department of Energy’s Oak Ridge National Laboratory used neutrons, isotopes and simulations to “see” the atomic structure of a saturated solution and found evidence supporting one of two competing hypotheses about how ions come

Biologists from Oak Ridge National Laboratory and the Smithsonian Environmental Research Center have confirmed that microorganisms called methanogens can transform mercury into the neurotoxin methylmercury with varying efficiency across species.


